Sent to CCL by: 'Close, David M.' [CLOSED]|[mail.etsu.edu]
The Gaussian manual has a good discussion of the ModRedundant command
under the keyword OPT.
You don't say just what you want to freeze. But you have the option of
freezing bond lengths, bond angles, and torsion angles. Just pick out
what you want to freeze, and type the frozen coordinates in a separate
section of the input file just after the geometry specifications. Add
the letter F (for frozen). For example if atom 4 is bonded to atom 5,
and you want this frozen at 1.50 A, type 5 4 1.50 F. There are useful
ways of freezing all bond lengths with * *, etc.
Another approach is to do a partial optimization (POPT) and separate the
geometries into two lists, Variables: and Constants:
Regards, Dave Close.
-----Original Message-----
> From: owner-chemistry+closedetsu.edu_._ccl.net
[mailto:owner-chemistry+closedetsu.edu_._ccl.net] On Behalf Of raj s k
cclselva]^[gmail.com
Sent: Tuesday, June 09, 2009 7:05 AM
To: Close, David M.
Subject: CCL:G: Help in freezing the coordinates in a Gaussian input
file.
Sent to CCL by: 'raj s k' [cclselva^-^gmail.com]
Hi,
Can anyone explain (with the syntax) how to freeze the coordinates of a
few atoms in a model in Gaussian input file ? I have about 30 atoms in a
model and about 24 atoms have be frozen before an optimization process.
It would be ideal if anyone can paste the text of an example Gaussian
input
file wherein few atomic coordinates are frozen.
Thanks
ks
-= This is automatically added to each message by the mailing script =-http://www.ccl.net/cgi-bin/ccl/send_ccl_messageSubscribe/Unsubscribe: Job: http://www.ccl.net/jobshttp://www.ccl.net/spammers.txt
-= This is automatically added to each message by the mailing script =-
To recover the email address of the author of the message, please change
the strange characters on the top line to the ^-^ sign. You can also
look up the X-Original-From: line in the mail header.
E-mail to subscribers: CHEMISTRY^-^ccl.net or use:
http://www.ccl.net/cgi-bin/ccl/send_ccl_message
E-mail to administrators: CHEMISTRY-REQUEST^-^ccl.net or use
http://www.ccl.net/cgi-bin/ccl/send_ccl_message
Subscribe/Unsubscribe:
http://www.ccl.net/chemistry/sub_unsub.shtml
Before posting, check wait time at: http://www.ccl.net
Job: http://www.ccl.net/jobs
Conferences: http://server.ccl.net/chemistry/announcements/conferences/
Search Messages: http://www.ccl.net/chemistry/searchccl/index.shtml
If your mail bounces from CCL with 5.7.1 error, check:
http://www.ccl.net/spammers.txt
RTFI: http://www.ccl.net/chemistry/aboutccl/instructions/
Oniom Input File Gaussian
Could Not Open Input File Artisan
ONIOM and Anion calculation in Gaussian? For two-layer ONIOM jobs, the format for this input line is. ESM_1.doc contains the Gaussian 09 input files and the optimized structures of the. Here is an input file for a two-layer ONIOM calculation using a DFT method for the high layer and Amber for the low layer. The molecule specification includes atom types (which are optional with UFF but required by Amber). Gaussian Tools. These tools were written for Gaussian. They work well for me; they may not work for you depending on your current Gaussian version and shell configuration. Route information of input file: g09-spin-contamination-after-annihilation. The rationale is that the output from that utility will not change between revisions of. Setting up ONIOM Calculations in Gaussian. There are several unique aspects to setting up an input file for a Gaussian ONIOM calculation: specifying the route, including layer assignments and other information with the molecule, and optionally specifying per-layer charges and spin multiplicities.
Input File Javascript
- This input file was created by GaussView. Note that it contains connectivity information for use with Geom =Connect. This job also illustrates the use of multiple charge and spin multiplicity values for ONIOM jobs.
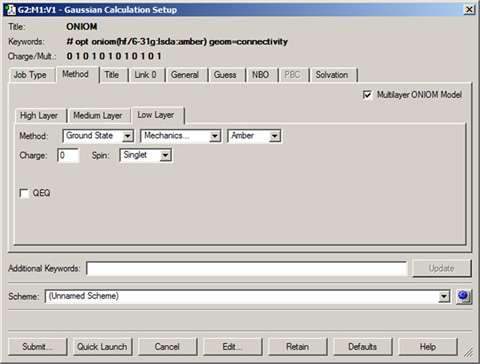
- The Gaussian Calculation Setup Window. The first step in producing a Gaussian input file is to build the desired molecule. The bond lengths, bond angles, and dihedral angles for the molecule will be used by GaussView to write a molecular structure for the calculation.
